Scientific Applications of MetaCyc |
|---|
Application 1: Metabolic Reconstruction for Sequenced Genomes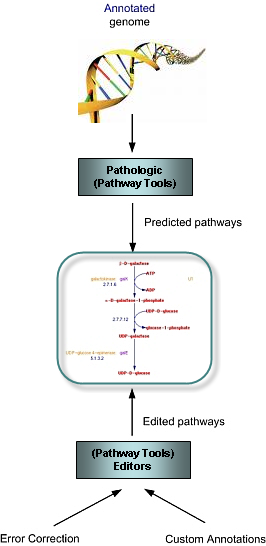
MetaCyc is an important aid for the computational reconstruction of metabolic networks from sequenced genomes. Using data stored in MetaCyc, pathway prediction software such as PathoLogic (a component of Pathway Tools) can be applied to an organism's complete, annotated [def] genomic sequence to infer its metabolic pathway complement. No need for pre-existing biochemistry: Computational pathway prediction is particularly helpful when little or no biochemistry is available for an organism of interest. The Pathway Tools software does not require biochemical knowledge about the organism of interest to predict its metabolic network.For this reason, computational prediction is particularly useful in cases where an organism is resistant to genetic manipulation, impeding its biochemical analysis, as demonstrated in the application of MetaCyc to understand sulfur assimilation in the extremophile Acidithiobacillus ferrooxidans based on a bioinformatics approach (Vald�s et al, 2003). Other examples include metabolic studies of the malaria pathogen Plasmodium falciparum 3D7 (Yeh et al, 2004), the prominent adult human gut symbiont Bacteroides thetaiotaomicron (Bjursell et al, 2006), the long-chain alkane degrading Geobacillus thermodenitrificans NG80-2, which was isolated from a deep-subsurface oil reservoir (Feng et al, 2007), the cellulolytic gliding bacterium Cytophaga hutchinsonii (Xie et al, 2007), the succinic acid producer Mannheimia succiniciproducens (Kim et al, 2007), the sap-feeding insect symbionts Sulcia muelleri and Baumannia cicadellinicola (McCutcheon and Moran, 2007), the pathogen Streptococcus pneumoniae (Aanensen et al, 2007), the butanol producer Clostridium acetobutylicum (Lee et al, 2008), the nitrogen fixer Azorhizobium caulinodans ORS571 (Lee et al, 2008), and many other organisms. A note of caution: Because these pathway complements are computationally predicted, they should be treated as first drafts until they have been reviewed manually. The Pathway Tools suite provides an extensive suite of editing tools to quickly fix errors and add custom annotations, including manually generated annotations, to the repository. Creating custom annotation of genomes: The downloadable version of Pathway Tools supports the editing of PGDBs [def] by users who want to generate and curate their own genome and pathway annotations. This approach provides a mechanism for capturing a laboratory's particular understanding of a research area. For example, a laboratory may want to alter an existing PGDB, such as provided by BioCyc, to annotate particulars region of a genome according to data generated within the laboratory. Examples of PGDBs that were created and curated by groups other that SRI is available at this web page. [top] |
Application 2: Support Metabolic Engineering and Drug Target DevelopmentMetabolic engineering is the targeted and purposeful alteration of an organism's metabolic pathways for research or industrial purposes. Metabolic pathways for e.g., the biosynthesis or degradation of substrates of commercial significance can be introduced or modified through genetic engineering to achieve alterations in pathway properties such as regulation and types of metabolites produced. Therefore, metabolic engineering is heavily dependent on the knowledge of all pathway variants and the availability of well-characterized enzymes and their genes as source materials for this process. MetaCyc provides a rich repository of both pathway variations and highly curated enzymes, and is routinely used in metabolic engineering projects as a reference (for example, see Drager et al 2007, Comparing various evolutionary algorithms on the parameter optimization of the valine and leucine biosynthesis in Corynebacterium glutamicum). In addition, researchers are increasingly using pathway information for both drug target development and pharmacogenomics, and the BioCyc PGDBs provide a key tool for this task (see Thorn et al 2007, Pathway-based Approaches to Pharmacogenomics). |
Application 3: Support Metabolomics ResearchMetaCyc provides a database of metabolites found in all domains of life that can aid metabolite identification. Metabolites are usually added to MetaCyc because they are substrates in a known metabolic reaction, or because they regulate the activity of a metabolic enzyme. MetaCyc also provides extensive data on metabolic reactions and enzymes that can provide insights regarding the potential routes of biosynthesis or catabolism of a given metabolite. Organism-specific metabolic reconstructions found in BioCyc provide a pathway context for interpretation of metabolomics data, via BioCyc metabolomics analysis tools for metabolite over-representation analysis, pathway-visualization of metabolomics data, and SmartTable transformations. |
Application 4: MetaCyc as an Encyclopedic Reference on Metabolic Pathways and EnzymesMetaCyc is used extensively as an online reference and in support of the teaching of:
|
Search
Genome
Metabolism
Desktop Application
Bring the power of BioCyc.org securely in-house, with the abilities to create and
edit PGDBs, perform metabolic modeling, and query/update using APIs.
Academic Licenses Commercial Licenses